publications
publications by categories in reversed chronological order. generated by jekyll-scholar.
* denotes equal contribution
2026
-
 Nature Communications 2026Magnetic resonance identification tags for ultra-flexible electrodesEminhan Özil, Peter Gombkoto, Athina Apostolelli, Tansel Baran Yasar, Angeliki D Vavladeli, Markus Marks, Manabu Rohr-Fukuma, Wolfger Behrens, and Mehmet Fatih Yanik
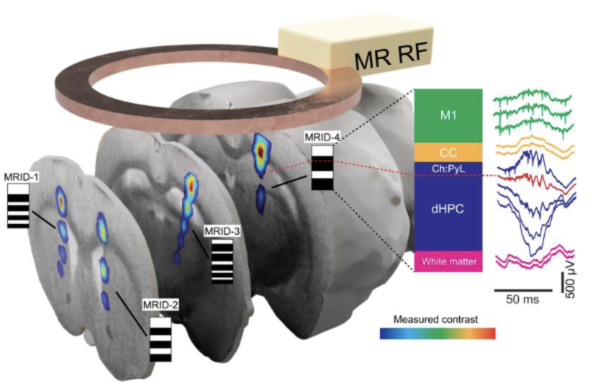
Nature Communications 2026Magnetic resonance identification tags for ultra-flexible electrodesEminhan Özil, Peter Gombkoto, Athina Apostolelli, Tansel Baran Yasar, Angeliki D Vavladeli, Markus Marks, Manabu Rohr-Fukuma, Wolfger Behrens, and Mehmet Fatih YanikUltra-flexible electrodes, due to their superior biocompatibility, are likely to lead the future of neuroprosthetics. However, identifying the precise positions of implanted high-density ultra-flexible electrodes in the brain for accurately assigning neural signals to specific structures remains a major challenge. To address this, we developed magnetic resonance identification (MRID)-tags. Each ultra-flexible electrode bundle carries an MRID-tag with unique barcode patterns visible in MRI (MRI-barcodes) for identification of the bundle. Individual bars in MRI-barcodes allow an accurate 3D reconstruction of the ultra-flexible electrode bundle’s trajectory in the brain and determine the anatomical positions of individual electrodes. We generate the MRI-barcodes by patterning superparamagnetic iron-oxide nanoparticles into electrode fibers (10 µm2) with dot-matrix nanoparticle coating technique. We chronically tested MRID-tagged ultra-flexible electrodes in vivo in the dorsal hippocampus of freely-moving rats, where distinct electrophysiological landmarks validated our electrode localization results. We were able to localize individual electrodes with a mean accuracy of 95 μm. MRID-tagged ultra-flexible electrodes demonstrated high long-term recording stability with mean single-unit signal-to-noise ratios as high as 20.
-
 ICLR 2026SAM 3: Segment Anything with ConceptsNicolas Carion, Laura Gustafson, Yuan-Ting Hu, Shoubhik Debnath, Ronghang Hu, Didac Suris, Chaitanya Ryali, Kalyan Vasudev Alwala, Haitham Khedr, Andrew Huang, Jie Lei, Tengyu Ma, Baishan Guo, Arpit Kalla, Markus Marks, and 23 more authors
ICLR 2026SAM 3: Segment Anything with ConceptsNicolas Carion, Laura Gustafson, Yuan-Ting Hu, Shoubhik Debnath, Ronghang Hu, Didac Suris, Chaitanya Ryali, Kalyan Vasudev Alwala, Haitham Khedr, Andrew Huang, Jie Lei, Tengyu Ma, Baishan Guo, Arpit Kalla, Markus Marks, and 23 more authorsWe present Segment Anything Model (SAM) 3, a unified model that detects, segments, and tracks objects in images and videos based on concept prompts, which we define as either short noun phrases (e.g., "yellow school bus"), image exemplars, or a combination of both. Promptable Concept Segmentation (PCS) takes such prompts and returns segmentation masks and unique identities for all matching object instances. To advance PCS, we build a scalable data engine that produces a high-quality dataset with 4M unique concept labels, including hard negatives, across images and videos. Our model consists of an image-level detector and a memory-based video tracker that share a single backbone. Recognition and localization are decoupled with a presence head, which boosts detection accuracy. SAM 3 doubles the accuracy of existing systems in both image and video PCS, and improves previous SAM capabilities on visual segmentation tasks. We open source SAM 3 along with our new Segment Anything with Concepts (SA-Co) benchmark for promptable concept segmentation.
-
 WACV 2026Diffusion-Based Action Recognition Generalizes to Untrained DomainsRogerio Guimaraes, Frank Xiao, Pietro Perona, and Markus Marks
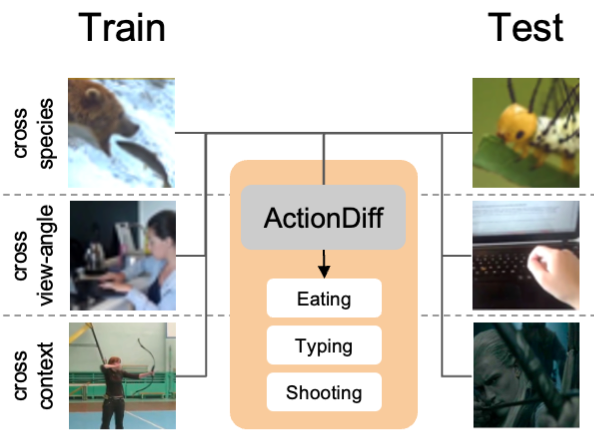
WACV 2026Diffusion-Based Action Recognition Generalizes to Untrained DomainsRogerio Guimaraes, Frank Xiao, Pietro Perona, and Markus MarksHumans can recognize the same actions despite large context and viewpoint variations, such as differences between species (walking in spiders vs. horses), viewpoints (egocentric vs. third-person), and contexts (real life vs movies). Current deep learning models struggle with such generalization. We propose using features generated by a Vision Diffusion Model (VDM), aggregated via a transformer, to achieve human-like action recognition across these challenging conditions. We find that generalization is enhanced by the use of a model conditioned on earlier timesteps of the diffusion process to highlight semantic information over pixel level details in the extracted features. We experimentally explore the generalization properties of our approach in classifying actions across animal species, across different viewing angles, and different recording contexts. Our model sets a new state-of-the-art across all three generalization benchmarks, bringing machine action recognition closer to human-like robustness.
-
 WACV 2026SAVeD: Learning to Denoise Low-SNR Video for Improved Downstream PerformanceSuzanne Stathatos, Michael Hobley, Pietro Perona*, and Markus Marks*
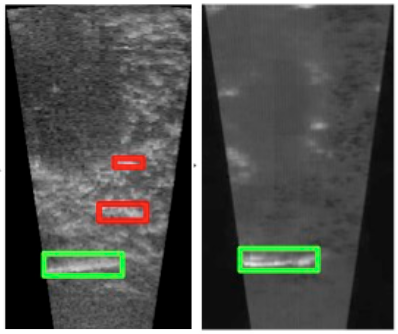
WACV 2026SAVeD: Learning to Denoise Low-SNR Video for Improved Downstream PerformanceSuzanne Stathatos, Michael Hobley, Pietro Perona*, and Markus Marks*Low signal-to-noise ratio videos – such as those from underwater sonar, ultrasound, and microscopy – pose significant challenges for computer vision models, particularly when paired clean imagery is unavailable. We present Spatiotemporal Augmentations and denoising in Video for Downstream Tasks (SAVeD), a novel self-supervised method that denoises low-SNR sensor videos using only raw noisy data. By leveraging distinctions between foreground and background motion and exaggerating objects with stronger motion signal, SAVeD enhances foreground object visibility and reduces background and camera noise without requiring clean video. SAVeD has a set of architectural optimizations that lead to faster throughput, training, and inference than existing deep learning methods. We also introduce a new denoising metric, FBD, which indicates foreground-background divergence for detection datasets without requiring clean imagery. Our approach achieves state-of-the-art results for classification, detection, tracking, and counting tasks, and it does so with fewer training resource requirements than existing deep-learning-based denoising methods.
2025
-
 Nature Methods 2025CellSAM: a foundation model for cell segmentationMarkus Marks*, Uriah Israel*, Rohit Dilip, Qilin Li, Morgan Schwartz, Elora Pradhan, Edward Pao, Shenyi Li, Alexander Pearson-Goulart, Pietro Perona, Georgia Gkioxari, Ross Barnowski, Yisong Yue, and David Ashley Van Valen
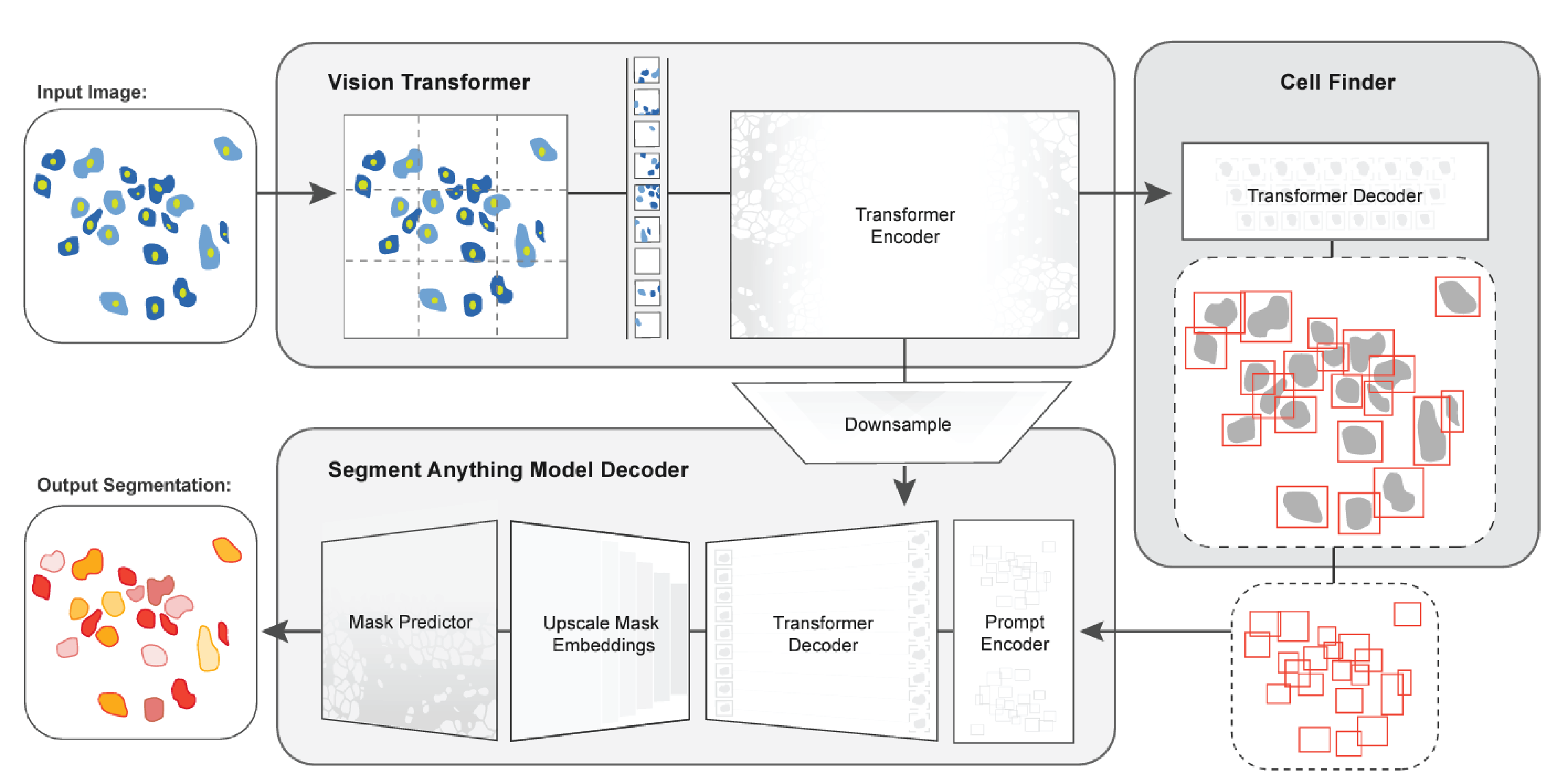
Nature Methods 2025CellSAM: a foundation model for cell segmentationMarkus Marks*, Uriah Israel*, Rohit Dilip, Qilin Li, Morgan Schwartz, Elora Pradhan, Edward Pao, Shenyi Li, Alexander Pearson-Goulart, Pietro Perona, Georgia Gkioxari, Ross Barnowski, Yisong Yue, and David Ashley Van ValenCells are a fundamental unit of biological organization, and identifying them in imaging data—cell segmentation—is a critical task for various cellular imaging experiments. Although deep learning methods have led to substantial progress on this problem, most models are specialist models that work well for specific domains but cannot be applied across domains or scale well with large amounts of data. Here we present CellSAM, a universal model for cell segmentation that generalizes across diverse cellular imaging data. CellSAM builds on top of the Segment Anything Model (SAM) by developing a prompt engineering approach for mask generation. We train an object detector, CellFinder, to automatically detect cells and prompt SAM to generate segmentations. We show that this approach allows a single model to achieve human-level performance for segmenting images of mammalian cells, yeast and bacteria collected across various imaging modalities. We show that CellSAM has strong zero-shot performance and can be improved with a few examples via few-shot learning. Additionally, we demonstrate how CellSAM can be applied across diverse bioimage analysis workflows.
-
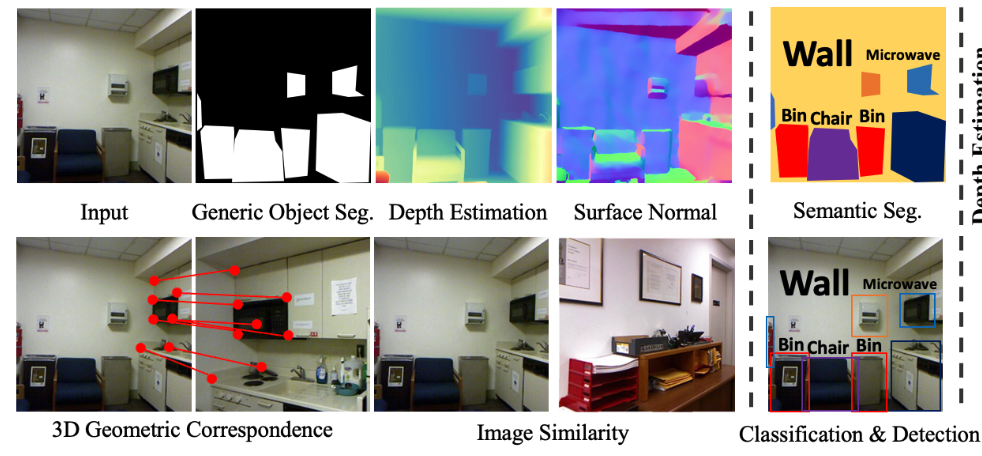 CVPR 2025Probing the Mid-level Vision Capabilities of Self-Supervised LearningXuweiyi Chen, Markus Marks, and Zezhou Cheng
CVPR 2025Probing the Mid-level Vision Capabilities of Self-Supervised LearningXuweiyi Chen, Markus Marks, and Zezhou ChengMid-level vision capabilities - such as generic object localization and 3D geometric understanding - are not only fundamental to human vision but are also crucial for many real-world applications of computer vision. These abilities emerge with minimal supervision during the early stages of human visual development. Despite their significance, current self-supervised learning (SSL) approaches are primarily designed and evaluated for high-level recognition tasks, leaving their mid-level vision capabilities largely unexamined. In this study, we introduce a suite of benchmark protocols to systematically assess mid-level vision capabilities and present a comprehensive, controlled evaluation of 22 prominent SSL models across 8 mid-level vision tasks. Our experiments reveal a weak correlation between mid-level and high-level task performance. We also identify several SSL methods with highly imbalanced performance across mid-level and high-level capabilities, as well as some that excel in both. Additionally, we investigate key factors contributing to mid-level vision performance, such as pretraining objectives and network architectures.
-
 WACV 2025Learning Keypoints for Multi-Agent Behavior Analysis using Self-SupervisionDaniel Khalil, Christina Liu, Pietro Perona, Jennifer Sun*, and Markus Marks*
WACV 2025Learning Keypoints for Multi-Agent Behavior Analysis using Self-SupervisionDaniel Khalil, Christina Liu, Pietro Perona, Jennifer Sun*, and Markus Marks*The study of social interactions and collective behaviors through multi-agent video analysis is crucial in biology. While self-supervised keypoint discovery has emerged as a promising solution to reduce the need for manual keypoint annotations, existing methods often struggle with videos containing multiple interacting agents, especially those of the same species and color. To address this, we introduce B-KinD-multi, a novel approach that leverages pre-trained video segmentation models to guide keypoint discovery in multi-agent scenarios. This eliminates the need for time-consuming manual annotations on new experimental settings and organisms. Extensive evaluations demonstrate improved keypoint regression and downstream behavioral classification in videos of flies, mice, and rats. Furthermore, our method generalizes well to other species, including ants, bees, and humans, highlighting its potential for broad applications in automated keypoint annotation for multi-agent behavior analysis.
-
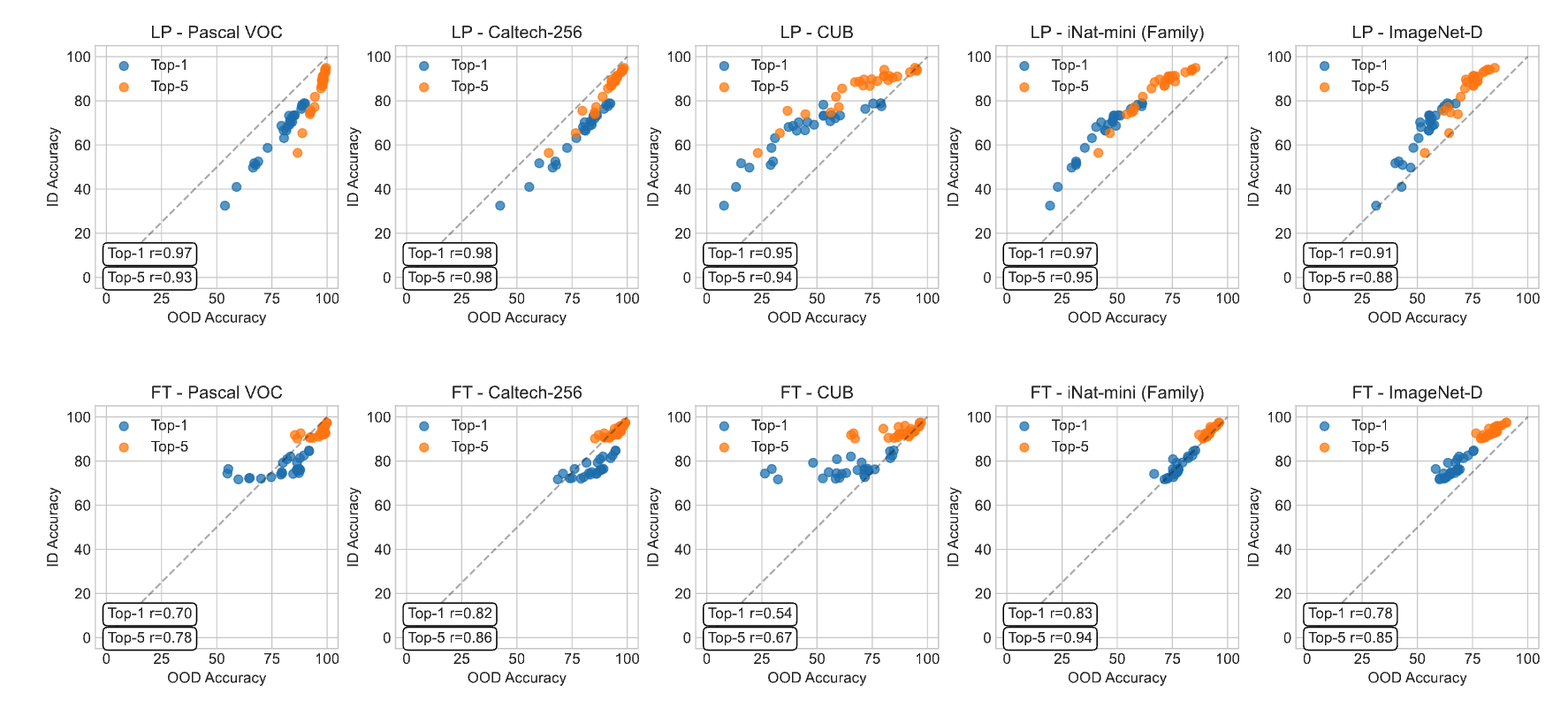 Int. Journal of Computer Vision 2025A Closer Look at Benchmarking Self-Supervised Pre-training with Image ClassificationMarkus Marks*, Manuel Knott*, Neehar Kondapaneni, Elijah Cole, Thijs Defraeye, Fernando Perez-Cruz, and Pietro Perona
Int. Journal of Computer Vision 2025A Closer Look at Benchmarking Self-Supervised Pre-training with Image ClassificationMarkus Marks*, Manuel Knott*, Neehar Kondapaneni, Elijah Cole, Thijs Defraeye, Fernando Perez-Cruz, and Pietro PeronaSelf-supervised learning (SSL) is a machine learning approach where the data itself provides supervision, eliminating the need for external labels. The model is forced to learn about the data structure or context by solving a pretext task. With SSL, models can learn from abundant and cheap unlabeled data, significantly reducing the cost of training models where labels are expensive or inaccessible. In Computer Vision, SSL is widely used as pre-training followed by a downstream task, such as supervised transfer, few-shot learning on smaller labeled data sets, and/or unsupervised clustering. Unfortunately, it is infeasible to evaluate SSL methods on all possible downstream tasks and objectively measure the quality of the learned representation. Instead, SSL methods are evaluated using in-domain evaluation protocols, such as fine-tuning, linear probing, and k-nearest neighbors (kNN). However, it is not well understood how well these evaluation protocols estimate the representation quality of a pre-trained model for different downstream tasks under different conditions, such as dataset, metric, and model architecture. We study how classification-based evaluation protocols for SSL correlate and how well they predict downstream performance on different dataset types. Our study includes eleven common image datasets and 26 models that were pre-trained with different SSL methods or have different model backbones. We find that in-domain linear/kNN probing protocols are, on average, the best general predictors for out-of-domain performance. We further investigate the importance of batch normalization and evaluate how robust correlations are for different kinds of dataset domain shifts.
2024
-
 ICLRw 2024Less is More: Discovering Concise Network ExplanationsNeehar Kondapaneni, Markus Marks, Oisin Mac Aodha, and Pietro Perona
ICLRw 2024Less is More: Discovering Concise Network ExplanationsNeehar Kondapaneni, Markus Marks, Oisin Mac Aodha, and Pietro PeronaWe introduce Discovering Conceptual Network Explanations (DCNE), a new approach for generating human-comprehensible visual explanations to enhance the interpretability of deep neural image classifiers. Our method automatically finds visual explanations that are critical for discriminating between classes. This is achieved by simultaneously optimizing three criteria: the explanations should be few, diverse, and human-interpretable. Our approach builds on the recently introduced Concept Relevance Propagation (CRP) explainability method. While CRP is effective at describing individual neuronal activations, it generates too many concepts, which impacts human comprehension. Instead, DCNE selects the few most important explanations. We introduce a new evaluation dataset centered on the challenging task of classifying birds, enabling us to compare the alignment of DCNE’s explanations to those of human expert-defined ones. Compared to existing eXplainable Artificial Intelligence (XAI) methods, DCNE has a desirable trade-off between conciseness and completeness when summarizing network explanations. It produces 1/30 of CRP’s explanations while only resulting in a slight reduction in explanation quality. DCNE represents a step forward in making neural network decisions accessible and interpretable to humans, providing a valuable tool for both researchers and practitioners in XAI and model alignment.
-
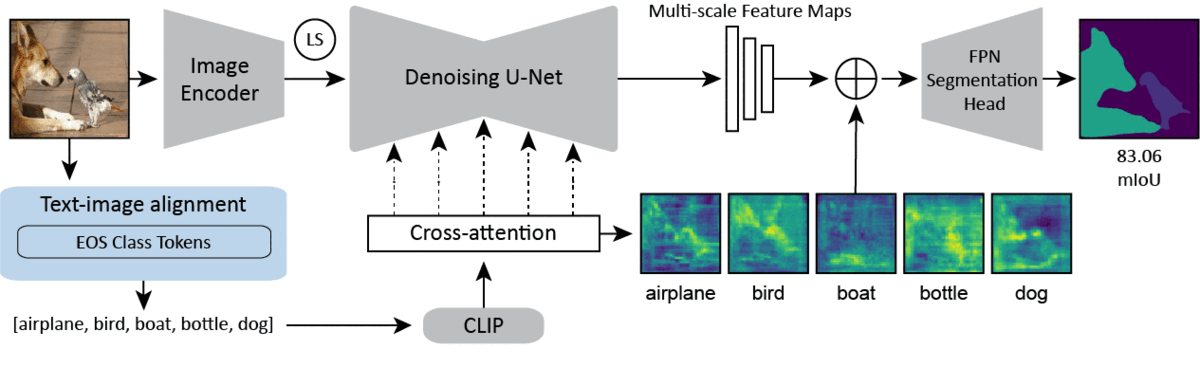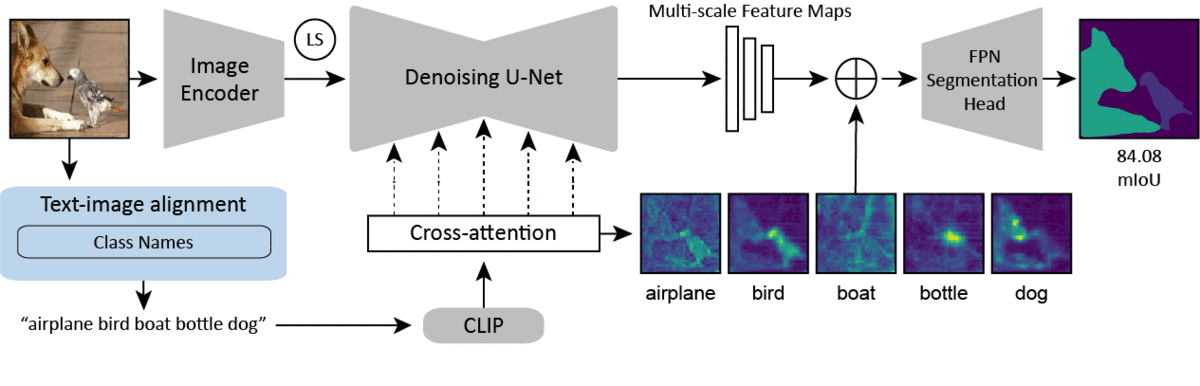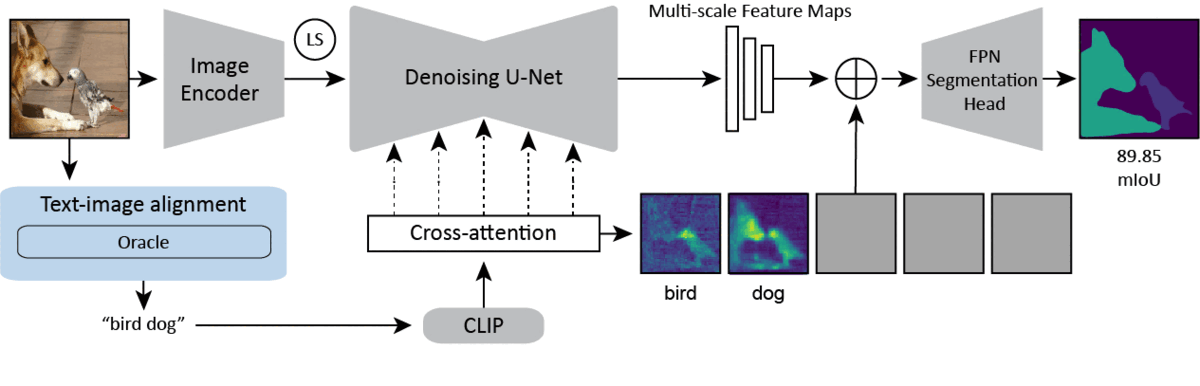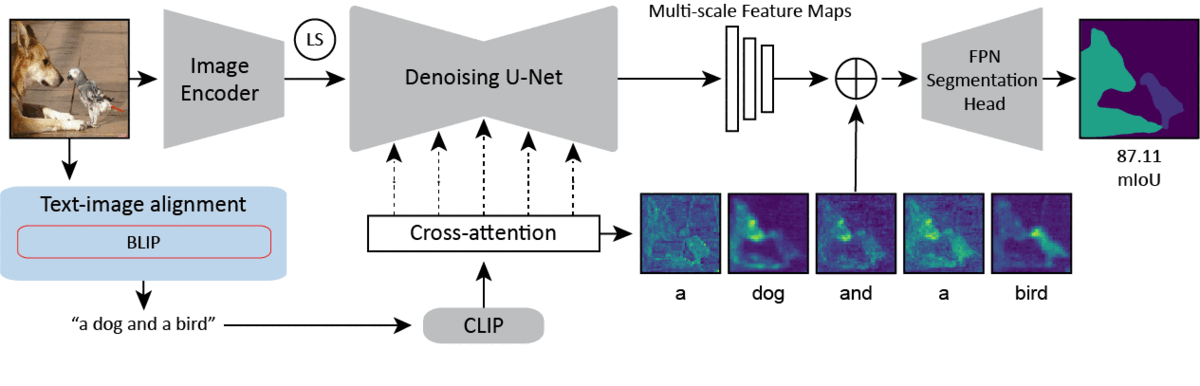 CVPR 2024Text-image Alignment for Diffusion-based PerceptionNeehar Kondapaneni*, Markus Marks*, Manuel Knott*, Rogério Guimarães, and Pietro Perona
CVPR 2024Text-image Alignment for Diffusion-based PerceptionNeehar Kondapaneni*, Markus Marks*, Manuel Knott*, Rogério Guimarães, and Pietro PeronaDiffusion models are generative models with impressive text-to-image synthesis capabilities and have spurred a new wave of creative methods for classical machine learning tasks. However, the best way to harness the perceptual knowledge of these generative models for visual tasks is still an open question. Specifically, it is unclear how to use the prompting interface when applying diffusion backbones to vision tasks. We find that automatically generated captions can improve text-image alignment and significantly enhance a model’s cross-attention maps, leading to better perceptual performance. Our approach improves upon the current state-of-the-art (SOTA) in diffusion-based semantic segmentation on ADE20K and the current overall SOTA for depth estimation on NYUv2. Furthermore, our method generalizes to the cross-domain setting. We use model personalization and caption modifications to align our model to the target domain and find improvements over unaligned baselines. Our cross-domain object detection model, trained on Pascal VOC, achieves SOTA results on Watercolor2K. Our cross-domain segmentation method, trained on Cityscapes, achieves SOTA results on Dark Zurich-val and Nighttime Driving.
2023
-
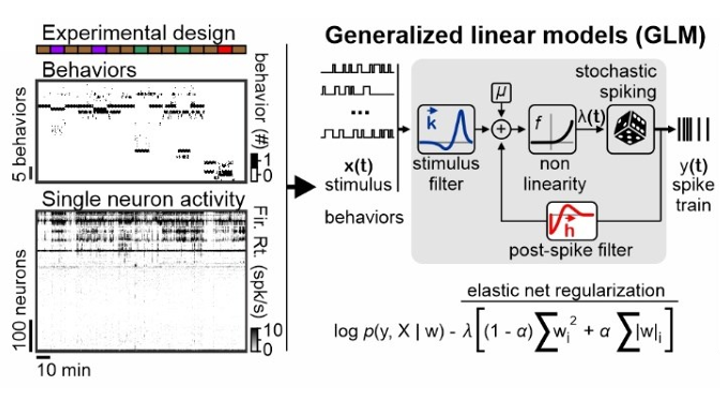 2023Distributed representations of innate behaviors in the hypothalamus do not predict specialized functional centersStefanos Stagkourakis, Giada Spigolon, Markus Marks, Michael Feyder, Joseph Kim, Pietro Perona, Marius Pachitariu, and David J Anderson
2023Distributed representations of innate behaviors in the hypothalamus do not predict specialized functional centersStefanos Stagkourakis, Giada Spigolon, Markus Marks, Michael Feyder, Joseph Kim, Pietro Perona, Marius Pachitariu, and David J AndersonArtificial activation of anatomically localized, genetically defined hypothalamic neuron populations is known to trigger distinct innate behaviors, suggesting a hypothalamic nucleus-centered organization of behavior control. To assess whether the encoding of behavior is similarly anatomically confined, we performed simultaneous neuron recordings across twenty hypothalamic regions in freely moving animals. Here we show that distinct but anatomically distributed neuron ensembles encode the social and fear behavior classes, primarily through mixed selectivity. While behavior class-encoding ensembles were spatially distributed, individual ensembles exhibited strong localization bias. Encoding models identified that behavior actions, but not motion-related variables, explained a large fraction of hypothalamic neuron activity variance. These results identify unexpected complexity in the hypothalamic encoding of instincts and provide a foundation for understanding the role of distributed neural representations in the expression of behaviors driven by hardwired circuits.
-
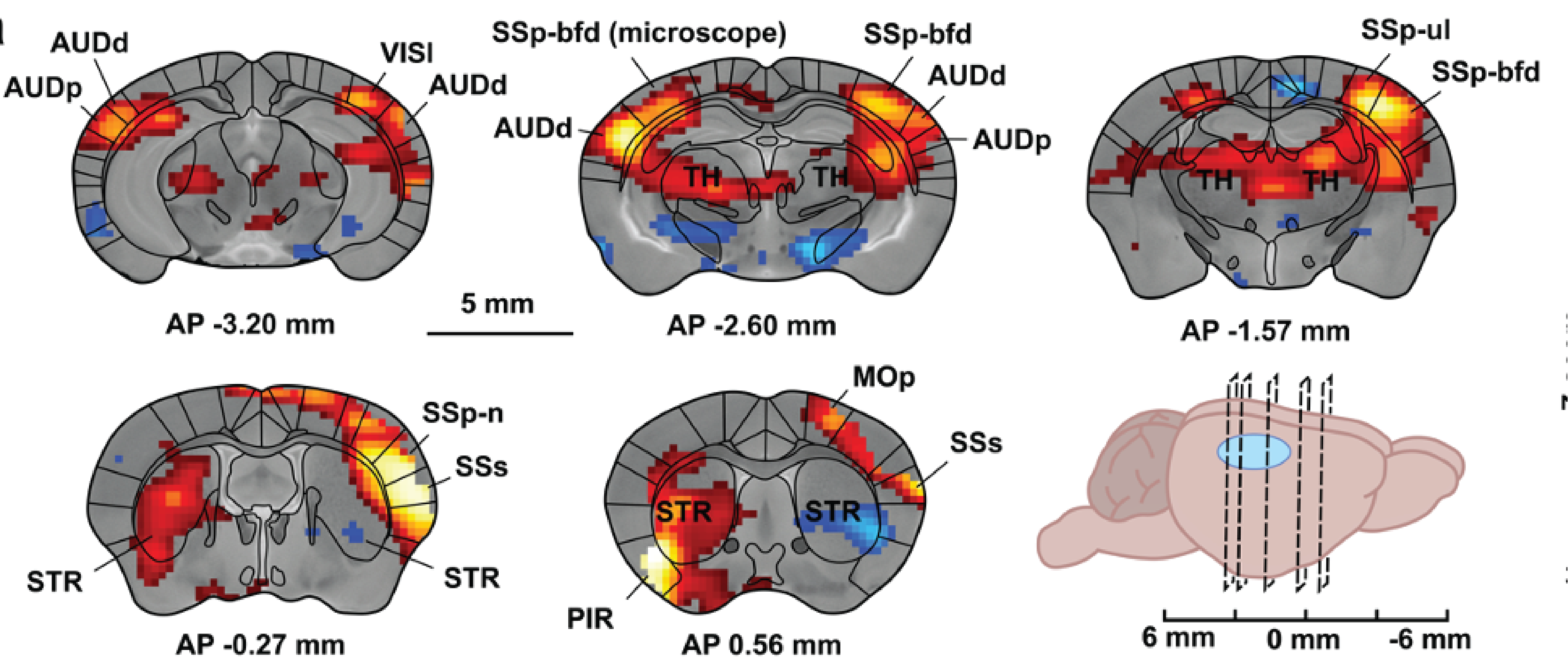 2023under review at Nature MethodsSimultaneous single-cell calcium imaging of neuronal population activity and brain-wide BOLD fMRIRik Lodevicus Elizabeth Maria Ubaghs, Roman Boehringer, Markus Marks, Helke Karen Hesse, Mehmet Fatih Yanik, Valerio Zerbi, and Benjamin Friedrich Grewe
2023under review at Nature MethodsSimultaneous single-cell calcium imaging of neuronal population activity and brain-wide BOLD fMRIRik Lodevicus Elizabeth Maria Ubaghs, Roman Boehringer, Markus Marks, Helke Karen Hesse, Mehmet Fatih Yanik, Valerio Zerbi, and Benjamin Friedrich GreweBlood Oxygen Level-Dependent (BOLD) functional Magnetic Resonance Imaging (fMRI) allows for non-invasive, indirect recordings of neural activity across the whole brain in both humans and animals. However, the relationship between the local neural population activity and the vascular activity is not completely understood. To investigate this relationship, we present a novel MRI compatible single-photon microscope capable of measuring cellular resolution Ca2+ activity of genetically defined neurons during whole-brain BOLD fMRI in awake behaving mice. Using this combined imaging approach, we found a difference in activity patterns between cells which was dependent on their location with respect to the vasculature. Notably, neurons near the vasculature showed pronounced negative activity during contralateral whisker movements at 3 Hz. In a second proof of concept experiment, we demonstrate the potential of recording both local neural activities, like those in the barrel field (SSp-bfd), and BOLD fMRI readings from interlinked brain regions. In sum, the presented technological advancement paves the way for studies examining the interplay between local brain circuits and overarching brain functions.
-
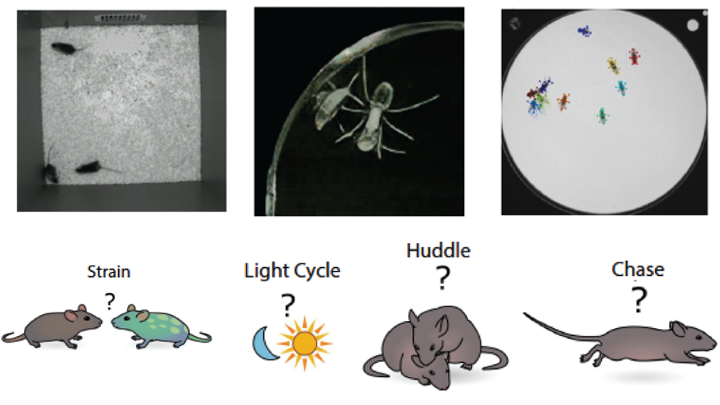 ICML 2023MABe22: A Multi-Species Multi-Task Benchmark for Learned Representations of BehaviorJennifer J. Sun*, Markus Marks*, Andrew Wesley Ulmer, Dipam Chakraborty, Brian Geuther, Edward Hayes, Heng Jia, Vivek Kumar, Sebastian Oleszko, Zachary Partridge, Milan Peelman, Alice Robie, Catherine E Schretter, Keith Sheppard, Chao Sun, and 8 more authors
ICML 2023MABe22: A Multi-Species Multi-Task Benchmark for Learned Representations of BehaviorJennifer J. Sun*, Markus Marks*, Andrew Wesley Ulmer, Dipam Chakraborty, Brian Geuther, Edward Hayes, Heng Jia, Vivek Kumar, Sebastian Oleszko, Zachary Partridge, Milan Peelman, Alice Robie, Catherine E Schretter, Keith Sheppard, Chao Sun, and 8 more authorsWe introduce MABe22, a large-scale, multi-agent video and trajectory benchmark to assess the quality of learned behavior representations. This dataset is collected from a variety of biology experiments, and includes triplets of interacting mice (4.7 million frames video+pose tracking data, 10 million frames pose only), symbiotic beetle-ant interactions (10 million frames video data), and groups of interacting flies (4.4 million frames of pose tracking data). Accompanying these data, we introduce a panel of real-life downstream analysis tasks to assess the quality of learned representations by evaluating how well they preserve information about the experimental conditions (e.g. strain, time of day, optogenetic stimulation) and animal behavior. We test multiple state-of-the-art self-supervised video and trajectory representation learning methods to demonstrate the use of our benchmark, revealing that methods developed using human action datasets do not fully translate to animal datasets.
2022
-
 Nature Machine Intelligence 2022Deep-learning-based identification, tracking, pose estimation and behaviour classification of interacting primates and mice in complex environmentsMarkus Marks, Qiuhan Jin, Oliver Sturman, Lukas Ziegler, Sepp Kollmorgen, Wolfger Behrens, Valerio Mante, Johannes Bohacek, and Mehmet Fatih Yanik
Nature Machine Intelligence 2022Deep-learning-based identification, tracking, pose estimation and behaviour classification of interacting primates and mice in complex environmentsMarkus Marks, Qiuhan Jin, Oliver Sturman, Lukas Ziegler, Sepp Kollmorgen, Wolfger Behrens, Valerio Mante, Johannes Bohacek, and Mehmet Fatih YanikQuantification of behaviours of interest from video data is commonly used to study brain function, the effects of pharmacological interventions, and genetic alterations. Existing approaches lack the capability to analyse the behaviour of groups of animals in complex environments. We present a novel deep learning architecture for classifying individual and social animal behaviour—even in complex environments directly from raw video frames—that requires no intervention after initial human supervision. Our behavioural classifier is embedded in a pipeline (SIPEC) that performs segmentation, identification, pose-estimation and classification of complex behaviour, outperforming the state of the art. SIPEC successfully recognizes multiple behaviours of freely moving individual mice as well as socially interacting non-human primates in three dimensions, using data only from simple mono-vision cameras in home-cage set-ups.
-
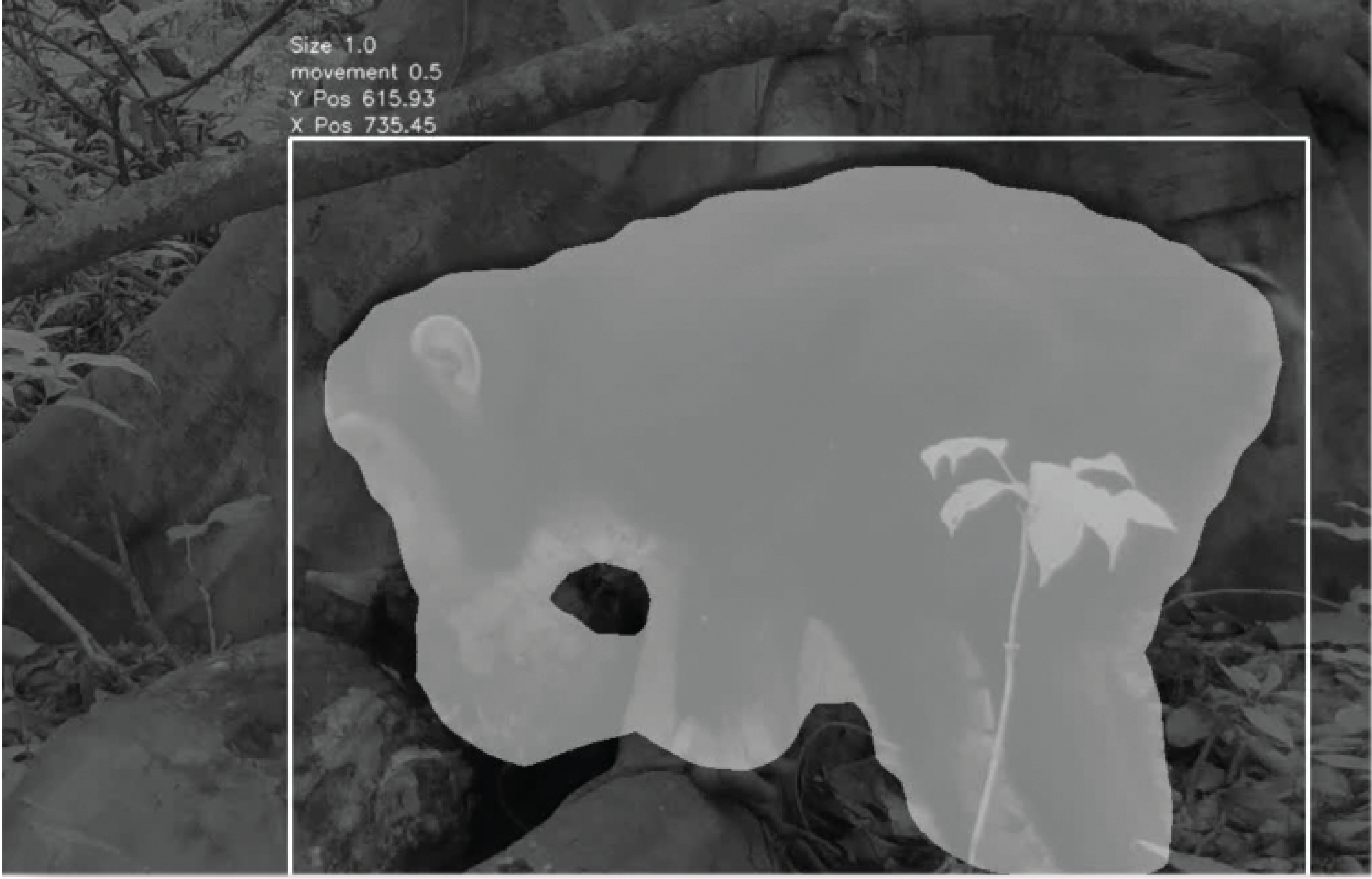 Journal of Animal Ecology 2022A multicomponent approach to studying cultural propensities during foraging in the wildKelly Ray Mannion, Elizabeth F Ballare, Markus Marks, and Thibaud Gruber
Journal of Animal Ecology 2022A multicomponent approach to studying cultural propensities during foraging in the wildKelly Ray Mannion, Elizabeth F Ballare, Markus Marks, and Thibaud GruberDetermining the cultural propensities or cultural behaviours of a species during foraging entails an investigation of underlying drivers and motivations. In this article, we propose a multi-component approach involving behaviour, ecology, and physiology to accelerate the study of cultural propensities in the wild. We propose as the first component the use of field experiments that simulate natural contexts, such as foraging behaviours and tool use opportunities, to explore social learning and cultural tendencies in a variety of species. To further accelerate this component, we discuss and advocate for the use of modern machine learning video analysis tools. In conjunction, we examine non-invasive methods to measure ecological influences on foraging such as phenology, fruit availability, dietary intake; and physiological influences such as stress, protein balance, energetics, and metabolism. To conclude, we highlight the benefits of combining ecological and physiological conditions with behavioural field experiments. This can be done across wild species, and provides the framework needed to test ecological hypotheses related to cultural behaviour.
- Int. Journal of CardiologyInt. Journal of Cardiology 2022Predicting cardiac remodeling after myocardial infarction with machine learning: are we there yet?Sebastian J Reinstadler, Clemens Dlaska, Martin Reindl, and Markus Marks
2021
-
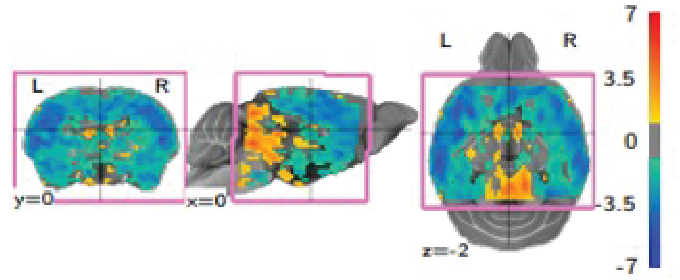 NeuroImage 2021An optimized registration workflow and standard geometric space for small animal brain imagingHorea-Ioan Ioanas, Markus Marks, Valerio Zerbi, Mehmet Fatih Yanik, and Markus Rudin
NeuroImage 2021An optimized registration workflow and standard geometric space for small animal brain imagingHorea-Ioan Ioanas, Markus Marks, Valerio Zerbi, Mehmet Fatih Yanik, and Markus RudinThe reliability of scientific results critically depends on reproducible and transparent data processing. Cross-subject and cross-study comparability of imaging data in general, and magnetic resonance imaging (MRI) data in particular, is contingent on the quality of registration to a standard reference space. In small animal MRI this is not adequately provided by currently used processing workflows, which utilize high-level scripts optimized for human data, and adapt animal data to fit the scripts, rather than vice-versa. In this fully reproducible article we showcase a generic workflow optimized for the mouse brain, alongside a standard reference space suited to harmonize data between analysis and operation. We introduce four separate metrics for automated quality control (QC), and a visualization method to aid operator inspection. Benchmarking this workflow against common legacy practices reveals that it performs more consistently, better preserves variance across subjects while minimizing variance across sessions, and improves both volume and smoothness conservation RMSE approximately 2-fold. We propose this open source workflow and the QC metrics as a new standard for small animal MRI registration.
2020
-
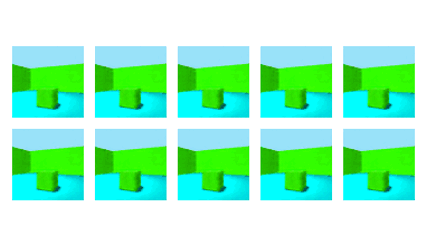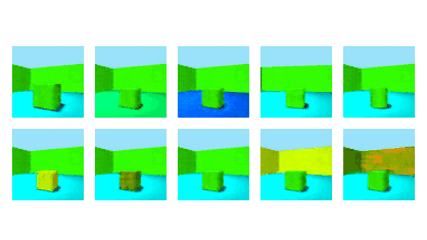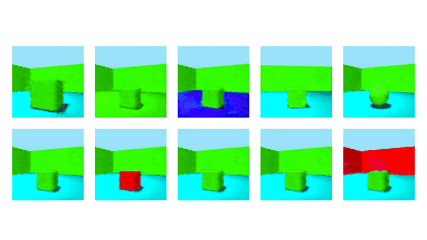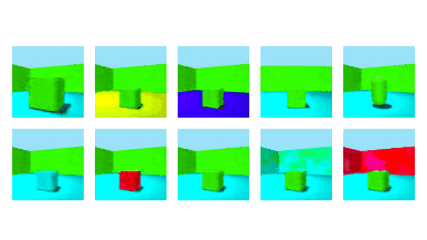 NeurIPS 2020Robust Disentanglement of a Few Factors at a Time using rPU-VAEBenjamin Estermann*, Markus Marks*, and Mehmet Fatih Yanik
NeurIPS 2020Robust Disentanglement of a Few Factors at a Time using rPU-VAEBenjamin Estermann*, Markus Marks*, and Mehmet Fatih YanikDisentanglement is at the forefront of unsupervised learning, as disentangled representations of data improve generalization, interpretability, and performance in downstream tasks. Current unsupervised approaches remain inapplicable for real-world datasets since they are highly variable in their performance and fail to reach levels of disentanglement of (semi-)supervised approaches. We introduce population-based training (PBT) for improving consistency in training variational autoencoders (VAEs) and demonstrate the validity of this approach in a supervised setting (PBT-VAE). We then use Unsupervised Disentanglement Ranking (UDR) as an unsupervised heuristic to score models in our PBT-VAE training and show how models trained this way tend to consistently disentangle only a subset of the generative factors. Building on top of this observation we introduce the recursive rPU-VAE approach. We train the model until convergence, remove the learned factors from the dataset and reiterate. In doing so, we can label subsets of the dataset with the learned factors and consecutively use these labels to train one model that fully disentangles the whole dataset. With this approach, we show striking improvement in state-of-the-art unsupervised disentanglement performance and robustness across multiple datasets and metrics.
-
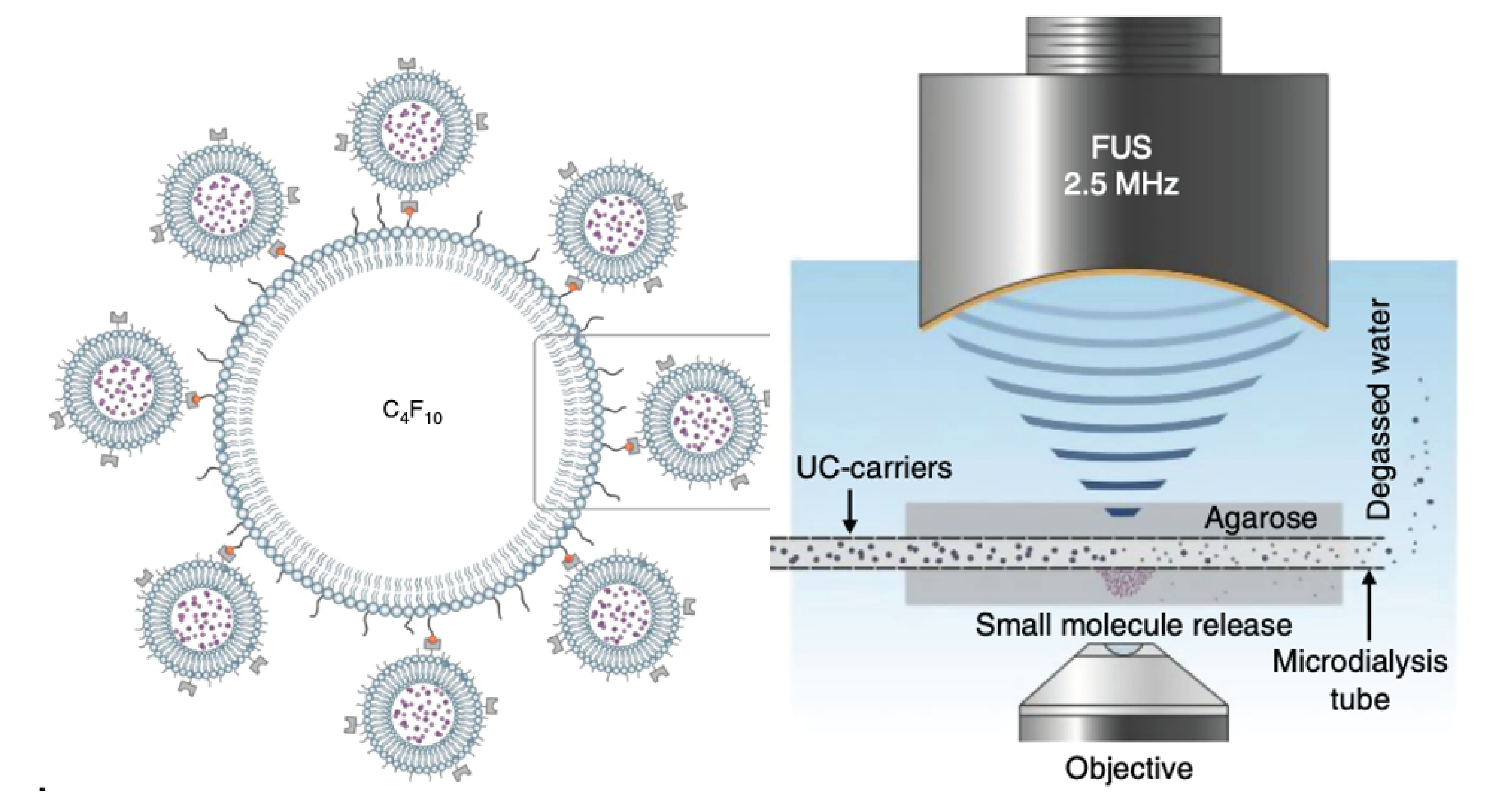 Nature Communications 2020Non-invasive molecularly-specific millimeter-resolution manipulation of brain circuits by ultrasound-mediated aggregation and uncaging of drug carriersMehmet S Ozdas, Aagam S Shah, Paul M Johnson, Nisheet Patel, Markus Marks, Tansel Baran Yasar, Urs Stalder, Laurent Bigler, Wolfger Behrens, Shashank R Sirsi, and others
Nature Communications 2020Non-invasive molecularly-specific millimeter-resolution manipulation of brain circuits by ultrasound-mediated aggregation and uncaging of drug carriersMehmet S Ozdas, Aagam S Shah, Paul M Johnson, Nisheet Patel, Markus Marks, Tansel Baran Yasar, Urs Stalder, Laurent Bigler, Wolfger Behrens, Shashank R Sirsi, and othersNon-invasive, molecularly-specific, focal modulation of brain circuits with low off-target effects can lead to breakthroughs in treatments of brain disorders. We systemically inject engineered ultrasound-controllable drug carriers and subsequently apply a novel two-component Aggregation and Uncaging Focused Ultrasound Sequence (AU-FUS) at the desired targets inside the brain. The first sequence aggregates drug carriers with millimeter-precision by orders of magnitude. The second sequence uncages the carrier’s cargo locally to achieve high target specificity without compromising the blood-brain barrier (BBB). Upon release from the carriers, drugs locally cross the intact BBB. We show circuit-specific manipulation of sensory signaling in motor cortex in rats by locally concentrating and releasing a GABAA receptor agonist from ultrasound-controlled carriers. Our approach uses orders of magnitude (1300x) less drug than is otherwise required by systemic injection and requires very low ultrasound pressures (20-fold below FDA safety limits for diagnostic imaging). We show that the BBB remains intact using passive cavitation detection (PCD), MRI-contrast agents and, importantly, also by sensitive fluorescent dye extravasation and immunohistochemistry.
-
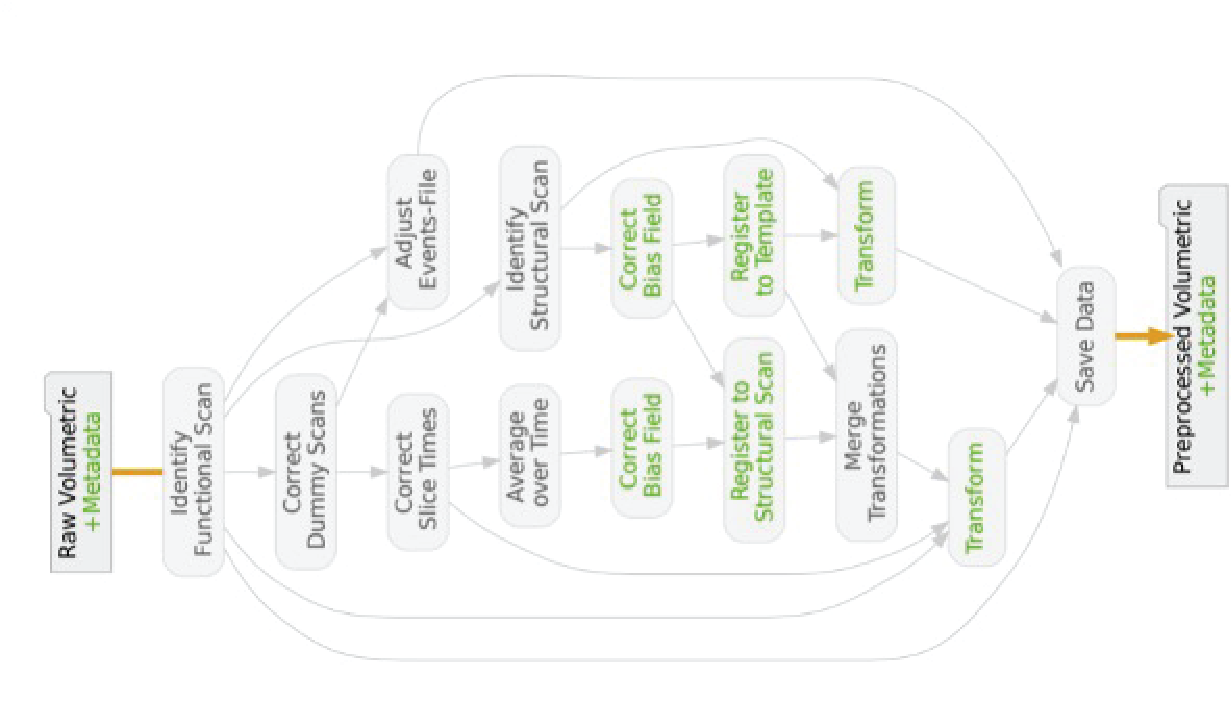 Frontiers in Neuroinformatics 2020An automated open-source workflow for standards-compliant integration of small animal magnetic resonance imaging dataHorea-Ioan Ioanas, Markus Marks, Clément M Garin, Marc Dhenain, Mehmet Fatih Yanik, and Markus Rudin
Frontiers in Neuroinformatics 2020An automated open-source workflow for standards-compliant integration of small animal magnetic resonance imaging dataHorea-Ioan Ioanas, Markus Marks, Clément M Garin, Marc Dhenain, Mehmet Fatih Yanik, and Markus RudinLarge-scale research integration is contingent on seamless access to data in standardized formats. Standards enable researchers to understand external experiment structures, pool results, and apply homogeneous preprocessing and analysis workflows. Particularly, they facilitate these features without the need for numerous potentially confounding compatibility add-ons. In small animal magnetic resonance imaging, an overwhelming proportion of data is acquired via the ParaVision software of the Bruker Corporation. The original data structure is predominantly transparent, but fundamentally incompatible with modern pipelines. Additionally, it sources metadata from free-field operator input, which diverges strongly between laboratories and researchers. In this article we present an open-source workflow which automatically converts and reposits data from the ParaVision structure into the widely supported and openly documented Brain Imaging Data Structure (BIDS). Complementing this workflow we also present operator guidelines for appropriate ParaVision data input, and a programmatic walk-through detailing how preexisting scans with uninterpretable metadata records can easily be made compliant after the acquisition.